# install devtools if needed
if (!requireNamespace("devtools")) {install.packages("devtools")}
# install from GitHub
devtools::install_github("sebastian-lortz/sspLNIRT")Overview
The sspLNIRT package provides sample size planning tools
for item calibration under the Joint Hierarchical Model (JHM; van der
Linden, 2007), combining a 2-parameter normal ogive model for response
accuracy with a log-normal model for response times, estimated via Gibbs
sampling through the LNIRT package (Fox et al., 2023).
The core question is: What is the minimum sample size to achieve a desired accuracy of item parameter estimates, given the assumed data-generating process? Accuracy is defined as the root mean squared error (RMSE) of a target item parameter (e.g., discrimination ) falling below a user-specified threshold, averaged across items and Monte Carlo iterations.
For a detailed description of the method, simulation model, and precomputed design conditions, see the accompanying manuscript.
Step 1: Specify the data-generating process
The first step is to specify the assumed data-generating process. This includes the item and person parameter distributions that define the simulation model. The following example uses a 30-item test () with the following specifications:
- Mean item parameters: , , , , with standard deviations of , , , and , respectively.
- Item difficulty () and time intensity () correlate at ; all other item parameter correlations are zero.
- Mean residual variance on the log scale: , with zero variance across items.
- Person parameters follow a bivariate standard normal distribution with ability–speed correlation .
sim.mod.args <- list(
K = 30,
mu.person = c(0, 0),
mu.item = c(1, 0, .5, 1),
meanlog.sigma2 = log(0.6),
sdlog.sigma2 = 0,
cov.m.person = matrix(c(1, 0.4,
0.4, 1), ncol = 2, byrow = TRUE),
cov.m.item = matrix(c(1, 0, 0, 0,
0, 1, 0, 0.4,
0, 0, 1, 0,
0, 0.4, 0, 1), ncol = 4, byrow = TRUE),
sd.item = c(.2, 1, .2, .5),
cor2cov.item = TRUE
)Inspecting implied distributions
Before committing to a design, it is useful to inspect the implied response accuracy and response time distributions. This helps evaluate whether the assumed data-generating process produces plausible score and time distributions for the intended application.
At the item level, plot_RA() shows the distribution of
implied response probabilities across items, and plot_RT()
shows within-item response time distributions:
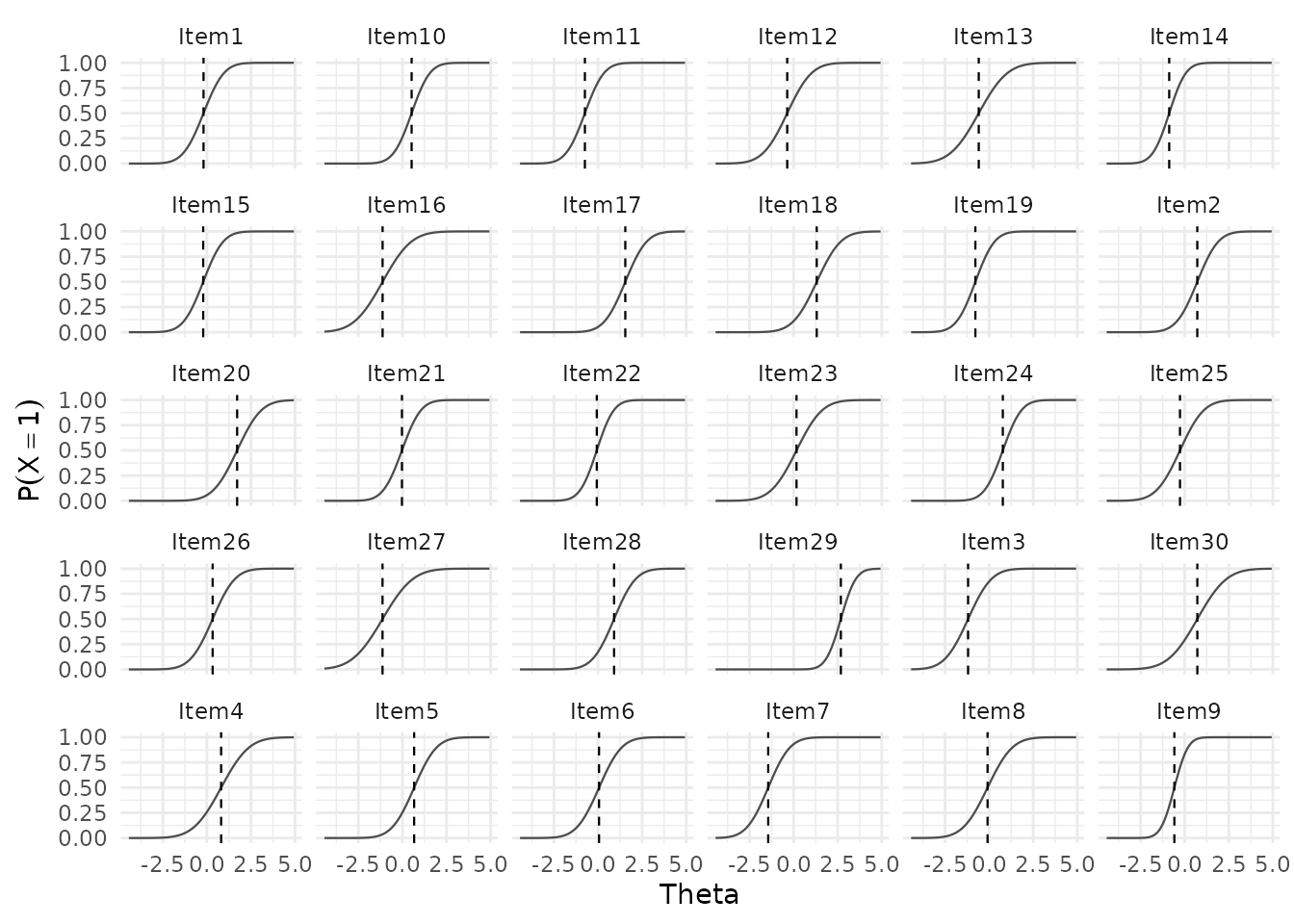
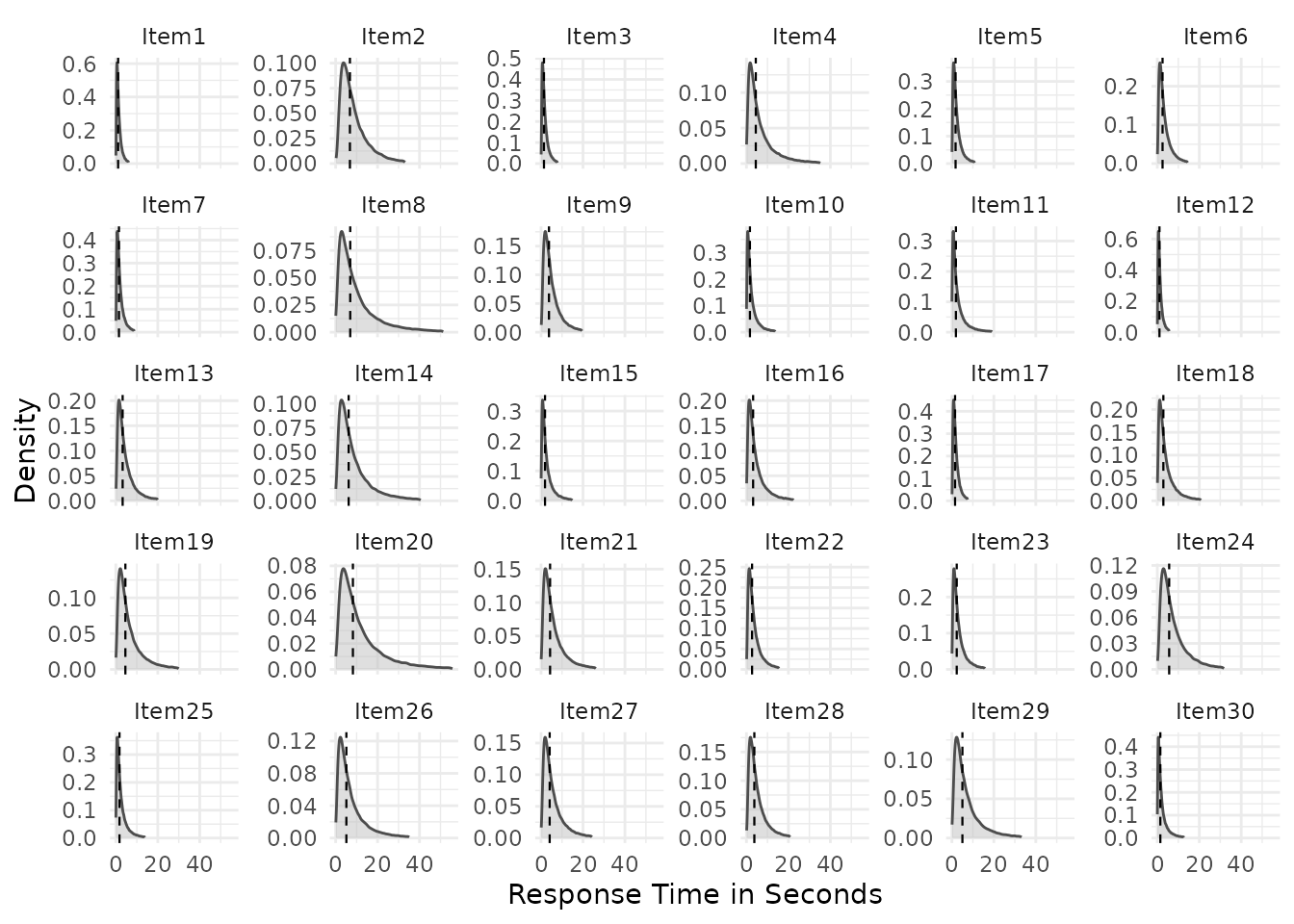
Additional views are available at the person level, showing total-correct score distributions and person-level (log-)RT distributions:
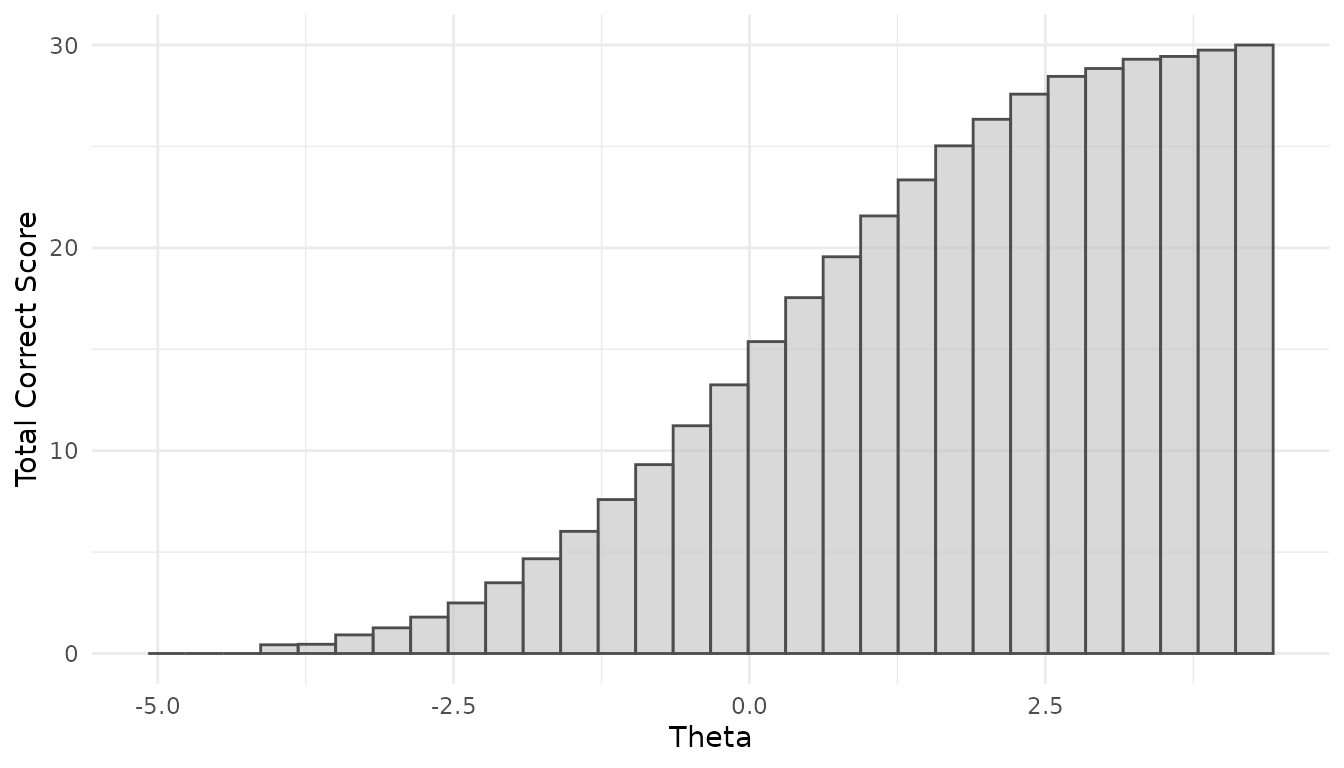
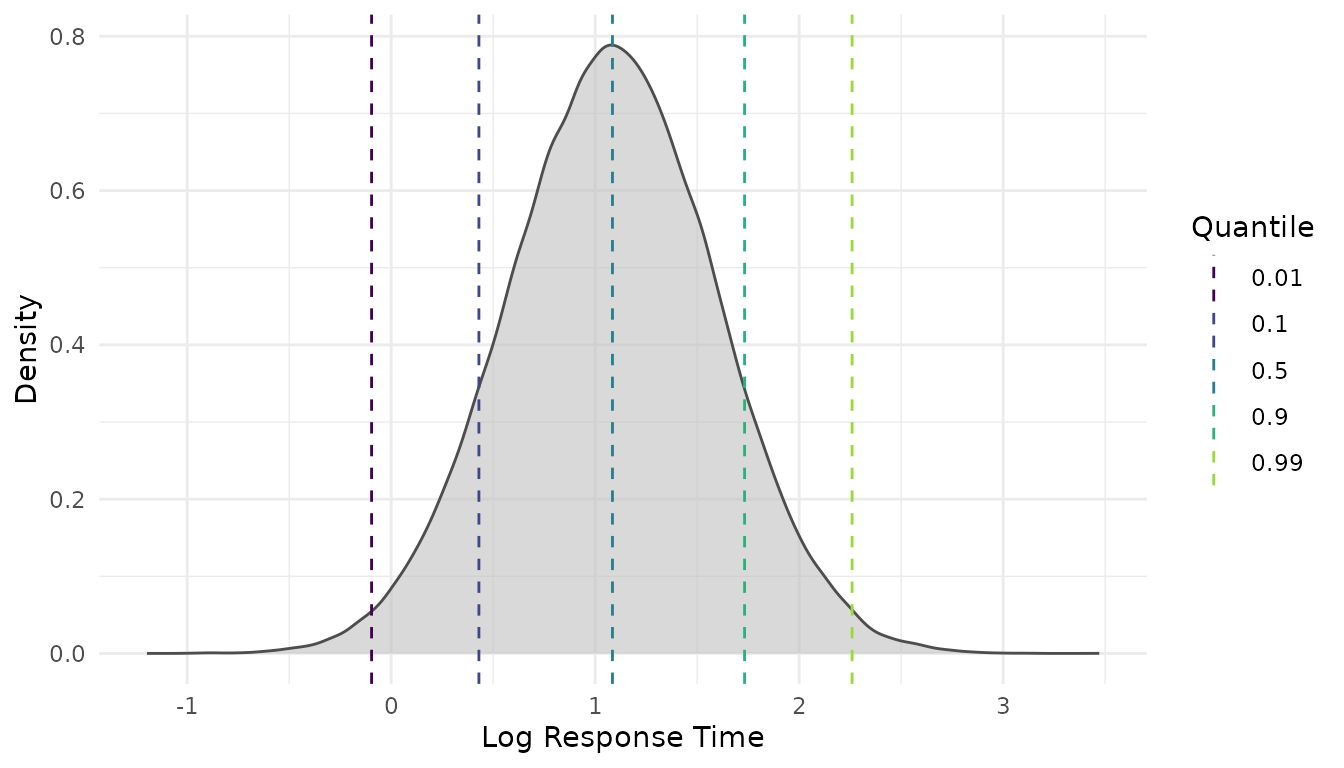
Both functions allow users to inspect how the simulated RA and RT distributions vary as a function of person ability and speed, respectively. This enables researchers to interpret the assumed data-generating process more accurately and facilitates making appropriate design choices.
Step 2: Determine the minimum sample size
Using precomputed results
The package comes with precomputed minimum sample sizes across a grid
of design conditions (see the manuscript for justifications on the
manipulated parameters). If the assumed design is covered by this grid,
results can be retrieved instantly without running any simulations. Use
available_configs() to check whether your specification
matches an available condition:
configs <- available_configs()
head(configs, 10)
#> thresh out.par K mu.alpha meanlog.sigma2 rho
#> 1 0.10 phi 30 1.4 -1.6094379 0.6
#> 2 0.20 phi 30 0.6 0.0000000 0.2
#> 3 0.05 alpha 10 1.0 -1.6094379 0.4
#> 4 0.20 beta 50 1.0 -1.6094379 0.4
#> 5 0.05 alpha 10 0.6 -0.5108256 0.6
#> 6 0.10 alpha 50 1.4 -0.5108256 0.6
#> 7 0.10 beta 30 1.4 0.0000000 0.4
#> 8 0.20 lambda 10 0.6 -0.5108256 0.4
#> 9 0.10 beta 10 1.4 -1.6094379 0.2
#> 10 0.10 lambda 10 0.6 0.0000000 0.6The columns correspond to the design factors that were varied:
thresh (RMSE threshold), out.par (target item
parameter), K (test length), mu.alpha (mean
item discrimination), meanlog.sigma2 (log-scale mean of
),
and rho (ability–speed correlation).
Optimizing for the minimum sample size
To optimize for the sample size one needs to specify the target
accuracy for the item parameters. Say that the target accuracies are
specified with
for the item parameters
,
respectively. These design and optimizer settings correspond to one of
the precomputed design conditions and can be retrieved from the
precomputed data with get_sspLNIRT():
result <- get_sspLNIRT(
thresh = c(0.1, 0.2, 0.05, 0.05),
out.par = c("alpha", "beta", "phi", "lambda"),
K = 30,
mu.alpha = 1,
meanlog.sigma2 = log(0.6),
rho = 0.4
)
summary(result$object)
#> ==================================================
#>
#> Call: optim_sample()
#>
#> Sample Size Optimization
#> --------------------------------------------------
#> Min Sample Size: 457
#> Optimizer Steps: 13
#>
#> Item Parameter RMSEs:
#> --------------------------------------------------
#> alpha beta phi lambda sigma2
#> RMSE 0.099416 0.112336 0.039268 0.041883 0.048074
#> MC SD 0.012777 0.022301 0.009195 0.009383 0.006420
#> Bias 0.000294 -0.008465 -0.000402 -0.001163 0.025169
#>
#> Person Parameter RMSEs:
#> --------------------------------------------------
#> theta zeta
#> RMSE 0.289977 0.258896
#> MC SD 0.013378 0.020407
#> Bias -0.006088 -0.001733
#> ---Because the precomputed results store each parameter separately,
get_sspLNIRT() looks up each
out.par/thresh pair individually and returns
the result for the bottleneck parameter — the one requiring the largest
sample size. The output reports the estimated minimum sample size
together with the estimated RMSE, Monte Carlo standard deviation of the
RMSE, and bias for all item and person parameters at
.
If the optimization stopped because the upper bound of the search
range was insufficient for any parameter (indicated by
N.min = "res.ub > thresh"), that parameter is
automatically selected as the bottleneck, signalling that the range
should be widened. Conversely, if all parameters already meet the
threshold at the lower bound
(N.min = "res.lb < thresh"), the first parameter’s
result is returned.
The function also works with scalar inputs. For example, to plan for item discrimination () only:
res_alpha <- get_sspLNIRT(
thresh = 0.1,
out.par = "alpha",
K = 30,
mu.alpha = 1,
meanlog.sigma2 = log(0.6),
rho = 0.4
)
summary(res_alpha$object)
#> ==================================================
#>
#> Call: optim_sample()
#>
#> Sample Size Optimization
#> --------------------------------------------------
#> Min Sample Size: 457
#> Optimizer Steps: 13
#>
#> Item Parameter RMSEs:
#> --------------------------------------------------
#> alpha beta phi lambda sigma2
#> RMSE 0.099416 0.112336 0.039268 0.041883 0.048074
#> MC SD 0.012777 0.022301 0.009195 0.009383 0.006420
#> Bias 0.000294 -0.008465 -0.000402 -0.001163 0.025169
#>
#> Person Parameter RMSEs:
#> --------------------------------------------------
#> theta zeta
#> RMSE 0.289977 0.258896
#> MC SD 0.013378 0.020407
#> Bias -0.006088 -0.001733
#> ---Using optim_sample() for custom designs
When the assumed design is not covered by the precomputed grid,
optim_sample() can be used to run the sample-size optimizer
directly. This uses a bisection algorithm that iteratively evaluates
comp_rmse() at candidate sample sizes until the RMSE of the
target parameter crosses the threshold. Each step runs a full Monte
Carlo simulation study with iter replications.
Like get_sspLNIRT(), optim_sample() accepts
vectors for thresh and out.par. The bisection
then requires all specified parameters to simultaneously fall below
their respective thresholds before narrowing the search interval, so the
resulting
satisfies all targets at once:
# Not run — computationally intensive (several hours)
library(future)
plan(multisession, workers = 4)
optim.args <- list(
thresh = c(0.1, 0.15, 0.05, 0.05),
range = c(50, 2000),
out.par = c("alpha", "beta", "phi", "lambda"),
iter = 200,
XG = 5000,
burnin = 20,
keep.err.dat = FALSE,
keep.rhat.dat = TRUE,
seed = 1234
)
res_custom <- do.call(optim_sample, c(sim.mod.args, optim.args))
summary(res_custom)For a single target parameter, scalar inputs work as before:
# Not run — computationally intensive
optim.args <- list(
thresh = 0.1,
range = c(50, 2000),
out.par = "alpha",
iter = 200,
XG = 5000,
burnin = 20,
keep.err.dat = FALSE,
keep.rhat.dat = TRUE,
seed = 1234
)
res_alpha <- do.call(optim_sample, c(sim.mod.args, optim.args))
summary(res_alpha)For a single evaluation at a fixed
(without optimization), use comp_rmse():
Step 3: Inspect and visualize
Power curve
The power curve shows how RMSE decreases with sample size, fitted as
a log-log regression through the optimization trace. The
out.par argument selects which parameter’s trace to plot,
the thresh argument plots a horizontal line for the
threshold:
plot_power_curve(result$object, out.par = "alpha", thresh = 0.1)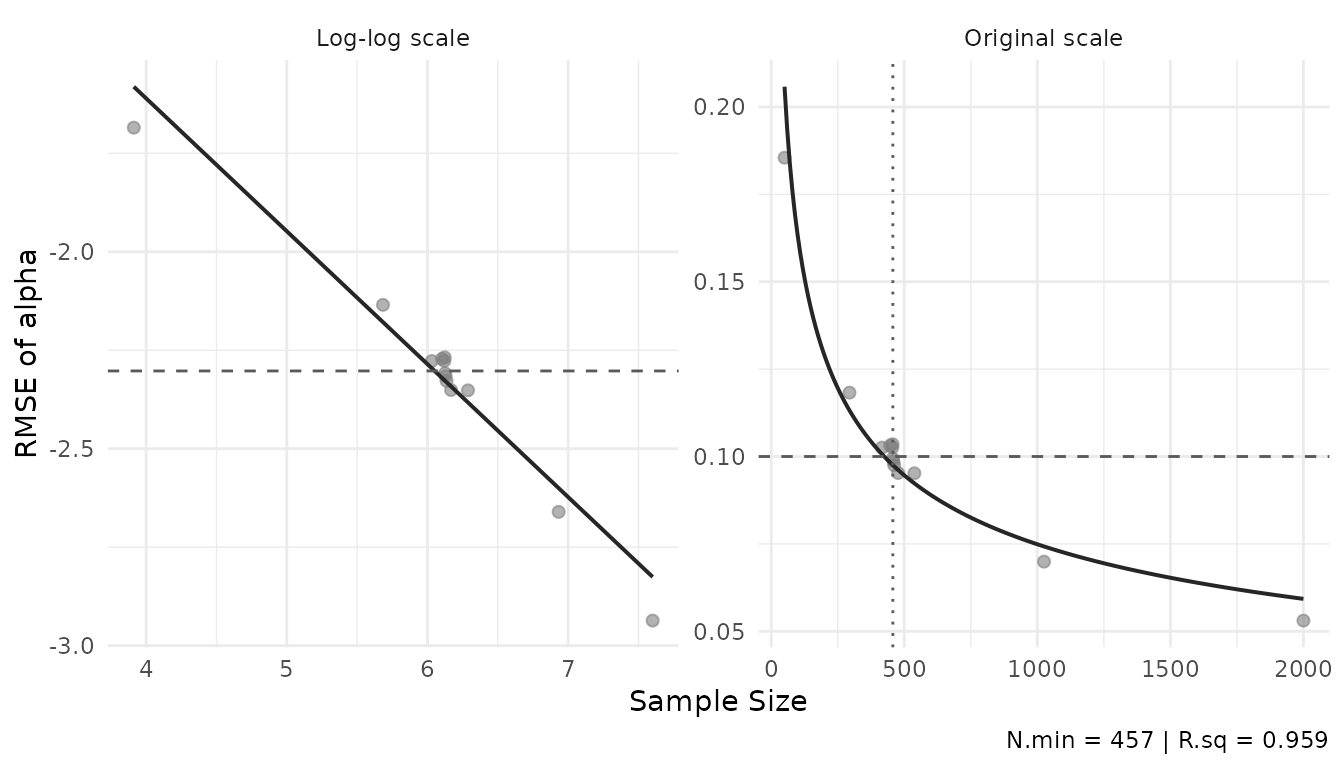
The left panel shows the log-log fit; the right panel the original scale. The dashed line marks the threshold, the dotted line the minimum .
Estimation accuracy
plot_estimation() shows how RMSE or bias varies across
the range of true parameter values at the minimum sample size
.
This is useful for identifying regions of the parameter space where
estimation is more or less precise — for instance, whether items with
extreme difficulty values are estimated less accurately.
plot_estimation(result$object, pars = "item", y.val = "rmse")
#> `geom_smooth()` using formula = 'y ~ x'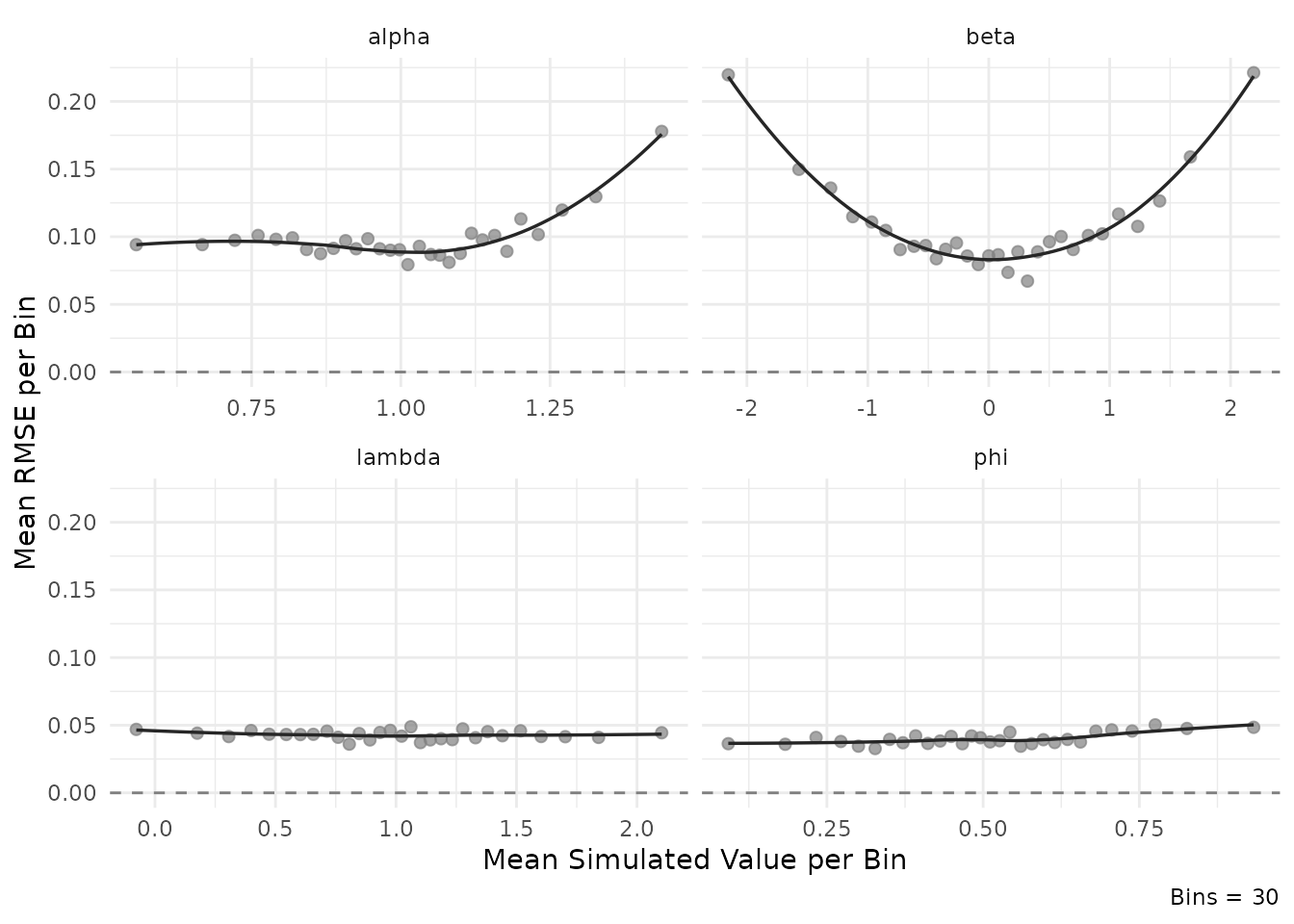
plot_estimation(result$object, pars = "person", y.val = "rmse")
#> `geom_smooth()` using formula = 'y ~ x'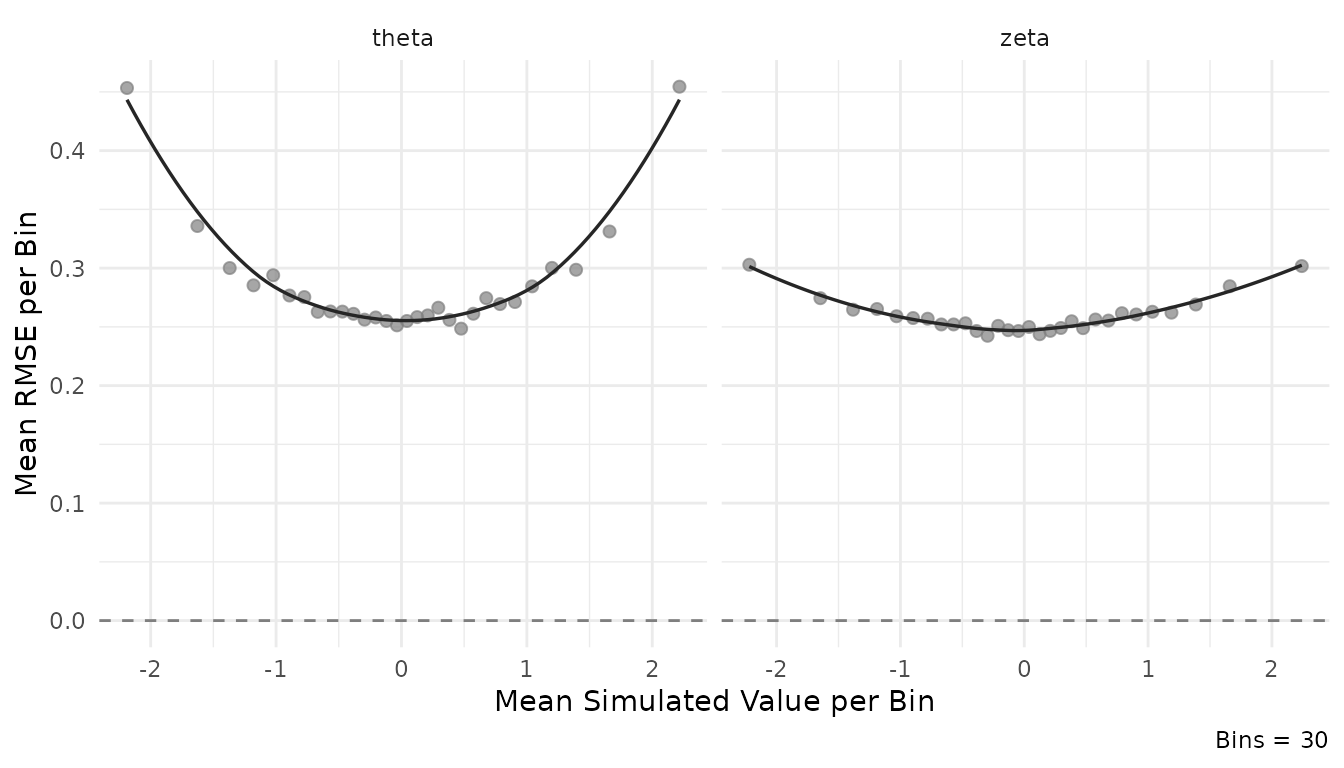
plot_estimation(result$object, pars = "item", y.val = "bias")
#> `geom_smooth()` using formula = 'y ~ x'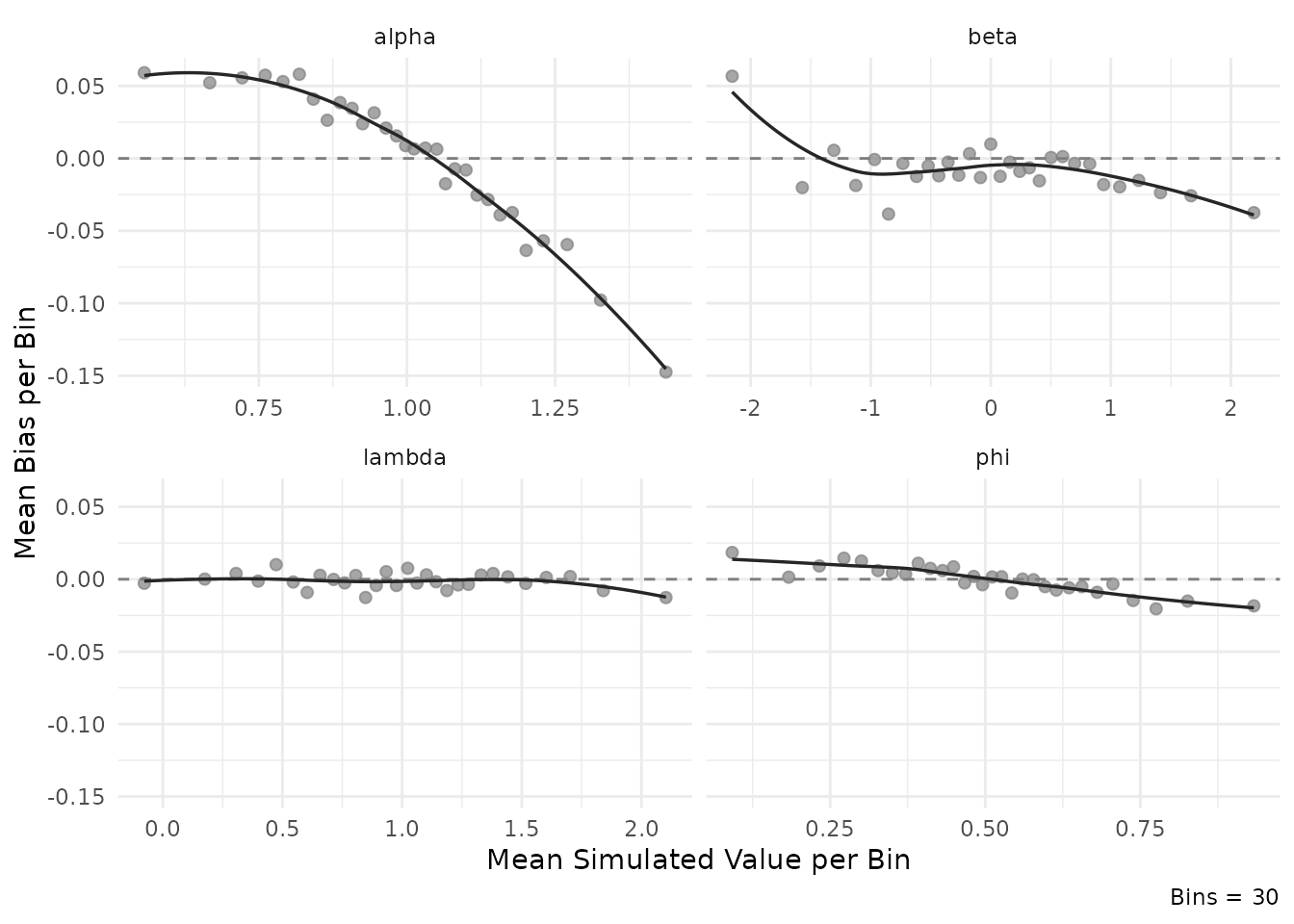
Design configuration
The full set of generating parameters can be inspected via the
design element of the output:
str(result$design)
#> List of 19
#> $ thresh : num 0.1
#> $ range : num [1:2] 50 2000
#> $ out.par : chr "alpha"
#> $ iter : num 100
#> $ K : num 30
#> $ mu.person : num [1:2] 0 0
#> $ mu.item : num [1:4] 1 0 0.5 1
#> $ meanlog.sigma2: num -0.511
#> $ cov.m.person : num [1:2, 1:2] 1 0.4 0.4 1
#> $ cov.m.item : num [1:4, 1:4] 1 0 0 0 0 1 0 0.4 0 0 ...
#> $ sd.item : num [1:4] 0.2 1 0.2 0.5
#> $ cor2cov.item : logi TRUE
#> $ sdlog.sigma2 : num 0
#> $ item.pars.m : NULL
#> $ XG : num 6000
#> $ burnin : num 20
#> $ seed : num 311263
#> $ rhat : num 1.05
#> $ keep.err.dat : logi FALSE
#> - attr(*, "class")= chr "sspLNIRT.design"Interactive Shiny application
The package includes a Shiny app for interactive exploration of precomputed results. Users specify design conditions via the interface, and the app retrieves the corresponding minimum sample size with summary statistics and visualizations. Alternatively, users can specify their own design condition, inspect the distribution visualizations, and download the function call for use in R.
run_app()References
Fox, J.-P., Klotzke, K., & Simsek, A. S. (2023). R-package LNIRT for joint modeling of response accuracy and times. PeerJ Computer Science, 9, e1232. https://doi.org/10.7717/peerj-cs.1232
van der Linden, W. J. (2007). A hierarchical framework for modeling speed and accuracy on test items. Psychometrika, 72(3), 287–308. https://doi.org/10.1007/s11336-006-1478-z